  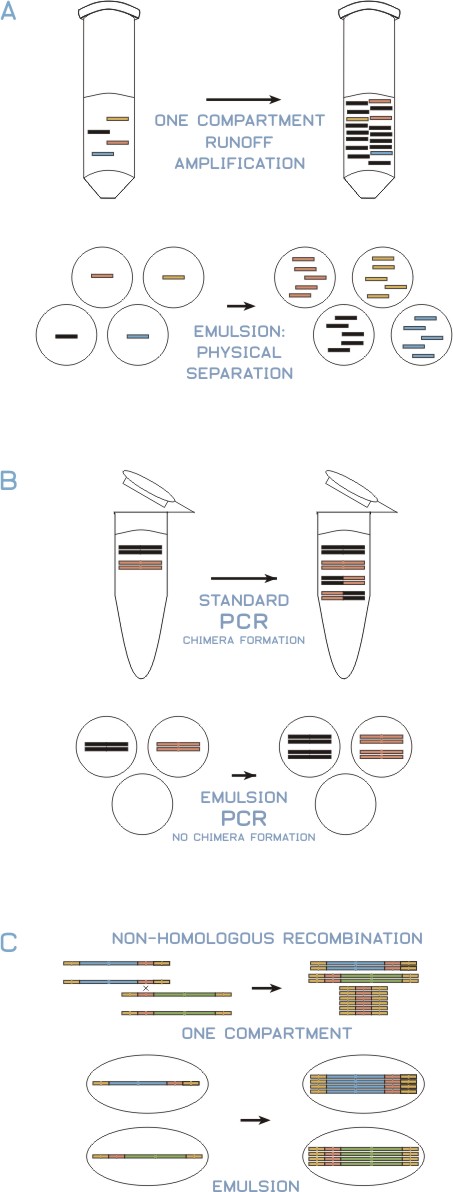
This problem with PCR may be overcome if each individual member of a library is amplified in a separate reaction. That is, in a random PCR environment, some DNA sequences are preferentially amplified at the expense of other sequences such that the amplified product does not represent the starting material. While random primed PCR can be easily engineered to amplify a plurality of nucleic acids in one reaction, this method is not preferred because the amplified library is not representative of the starting library. 
Other techniques, such as PCR, while fast and reliable, are unable to amplify a genome in a representative fashion. Current techniques for in vitro genome amplification involve laborious cloning and culturing protocols that have limited the utility of genomic sequencing. The starting material may be limited, for example, if the genome to be sequenced is from a trace quantity of pathogen or from a prenatal patient. If the starting material is limited, amplification of the initial DNA is necessary before genomic sequencing. Furthermore, the sequencing of a human genome would require about tens of millions of different sequencing reactions. Current sequencing technologies require millions of copies of nucleic acid per sequencing reaction. The ability to amplify a plurality of nucleic acid sequences, such as a genomic library or a cDNA library, is critical given the inefficiency of current methods of sequencing. The present invention is also directed to zero bead removal-a method of enriching for solid support containing amplified nucleic acids is also disclosed. The present invention relates to methods for amplifying nucleic acid templates from low copy number to quantities amenable for sequencing on a solid support such as a bead. 16, 2019 and is 6 KB in size, are hereby incorporated by reference in their entirety. The contents of the text file named “454L_008C07U_Sequence_Listing.txt”, which was created on Jan. INCORPORATION BY REFERENCE OF SEQUENCE LISTING 17, 2007, and “Methods Of Amplifying And Sequencing Nucleic Acids” filed on Jan. 
10/767,894, now abandoned, “Double Ended Sequencing” filed on Jan. patent applications: “Method For Preparing Single-Stranded DNA Libraries” filed on Jan. This application also incorporates by reference the following U.S. All patent and patent applications in this paragraph are hereby fully incorporated herein by reference. 28, 2004, which claims the benefit of priority to the following applications: U.S. 31, 2007, which is a continuation of U.S. 23, 2011, which is a continuation of U.S. 14, 2012, which is a continuation of U.S. 25, 2013, now abandoned, which is a continuation of U.S. 20, 2014, which is a continuation of U.S. 23, 2016, which is a continuation of U.S. Yu B (2014) Setting up next-generation sequencing in the medical laboratory.This application is a continuation of U.S.Yu B (2014) Setting up next-generation sequencing in the medical laboratory.Xuan J, Yu Y, Qing T et al (2013) Next-generation sequencing in the clinic: promises and challenges.Williams R, Peisajovich SG, Miller OJ et al (2006) Amplification of complex gene libraries by emulsion PCR.Meyerhans A, Vartanian JP, Wain-Hobson S (1990) DNA recombination during PCR.Dressman D, Yan H, Traverso G et al (2003) Transforming single DNA molecules into fluorescent magnetic particles for detection and enumeration of genetic variations.Nakano M, Komatsu J, Matsuura S et al (2003) Single-molecule PCR using water-in-oil emulsion.Buermans HP, den Dunnen JT (2014) Next generation sequencing technology: advances and applications.Mamanova L, Coffey AJ, Scott CE et al (2010) Target-enrichment strategies for next-generation sequencing.Gullapalli RR, Desai KV, Santana-Santos L et al (2012) Next generation sequencing in clinical medicine: challenges and lessons for pathology and biomedical informatics.Metzker ML (2010) Sequencing technologies – the next generation. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
